Technology
Ion channels are promising drug targets with diverse physiological functions and severe impact in diseases like autoimmune, cancer, pain and others. The high degree of structural similarity make ion channels a hard-to-hit target as it requires a multi-epitope binding for drug selectivity. Many small molecules and antibodies programs have failed due to structural deficits which lead to unselective binding. A broad evolutionary convergence as seen in mollusks, insects, vertebrates and even plants has resulted in peptides as potent ion channel blockers.
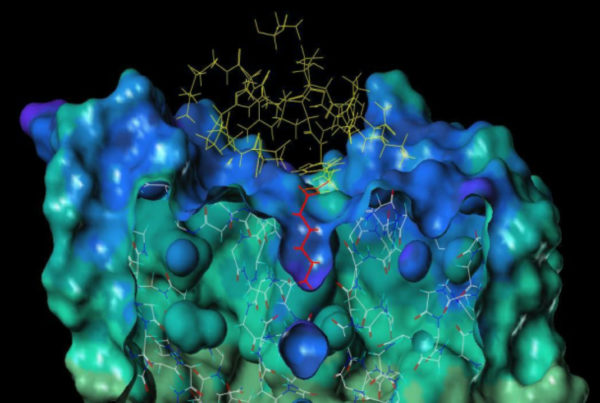
In recent years genomic data and x-ray crystallography have helped to understand the structural characteristics of ion channels and peptides and to develop selectION’s si-544 lead candidate for selective ion channel binding.
Peptide Structures
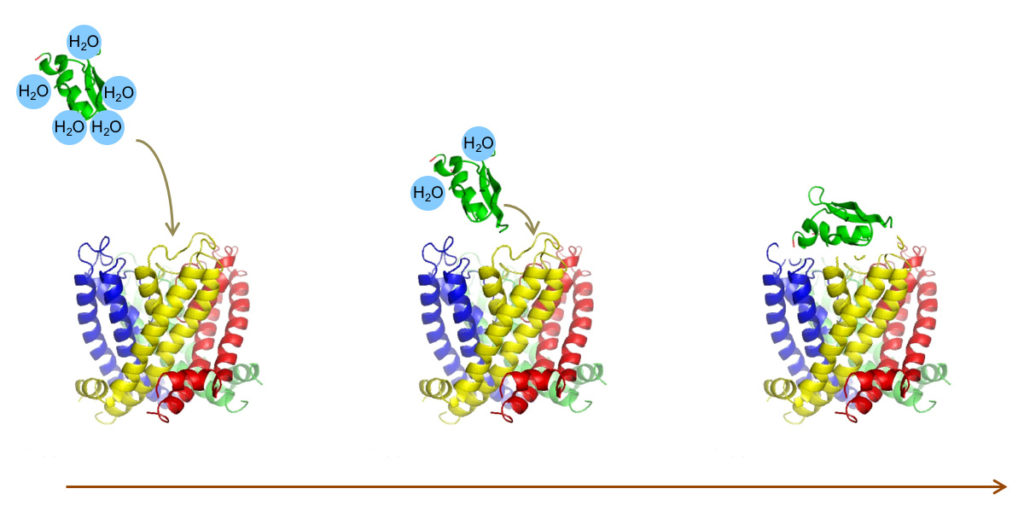
selectION uses peptide structures derived from evolutionary tuned toxins from venomous animals like scorpions or conesnails as blueprints for its screening platform. These molecules are highly constrained peptides typically between 20 and 40 amino acids in size and stabilized by an intramolecular scaffold of 2–4 disulfide bridges. A low molecular weight 2–4.5kDa combined with high water solubility make them ideal drug candidates. From a regulatory perspective peptides can be regarded as small molecules offering several advantages during the developmental process.

Peptide Selectivity
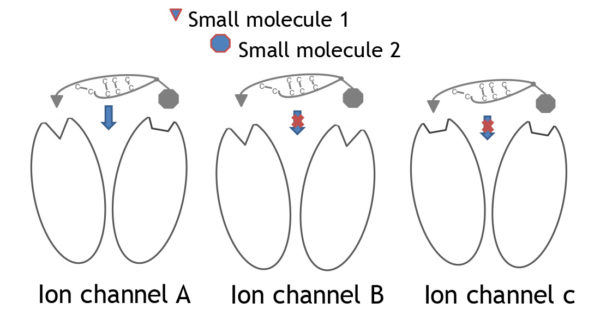
3 different ion channels (A, B, C) each with 2 subunits are shown. Each ion channel displays a unique combination of binding pockets that mediate selective binding to the channel. The drafted peptide needs to bind simultaneously to both binding pockets for full antagonistic action. Selective binding is only given with subtype A whereas the small molecules 1 and 2 can bind to at least 2 channel subtypes.
Screening Platform

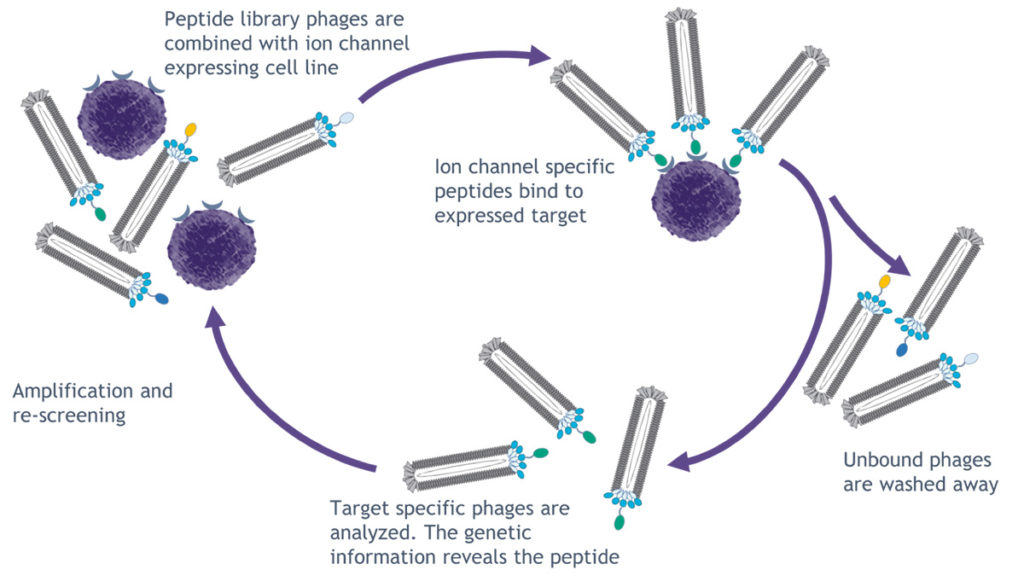
selectION has established a modified phage display platform to identify highly selective peptide binders for ion channels. Known ion channel binding peptides derived from animal venoms are employed as blueprints in the screening process. Based on these natural scaffolds new ion channel targeted peptide libraries are designed. These libraries are screened in multiple rounds of peptide selection on a phage display platform. Subsequently, binding phages are recovered and peptide sequences are determined via sequencing.
The technology can be implemented on every single ion channel target expressed in a cellular system combined with off-targets screens to reduce unselective peptides.